news
Understanding the Origins of Cancer: Scientists Investigate the Molecular Changes that Lead to Disease
Primary tabs
Written by Abby Vogel
Cancer is the most-feared of human diseases, often striking without warning and seemingly without identifiable cause. Decades into the nation’s war on cancer, we have learned that the disease is far more complex than we originally believed.
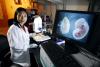
Kirill Lobachev, associate professor in the School of Biology, uses this gel documentation system to capture images of DNA on a gel. (Click image for high-resolution version. Credit: Gary Meek)
At the Georgia Institute of Technology, researchers are pursuing many different directions in their quest to understand how cancer arises. They are adding their findings to a deepening understanding of the complex molecular pathways that turn a normal cell into a malignant one. Ultimately, that knowledge may lead to new strategies for preventing cancer, new diagnostic techniques for finding it early – and to drugs and other agents that may provide cures.
This article describes Georgia Tech research into the origins of cancer including:
- How hormones fuel certain cancers;
- The potential role of non-mutational changes, called epigenetics;
- An integrated approach to studying ovarian cancer;
- Mechanisms for repairing double-strand DNA breaks;
- Predicting where DNA will break, and how often it will break; and
- Understanding the role of cell-signaling molecules such as sphingolipids.
This is the first in a series of three reports that will focus on cancer research at Georgia Tech. The other two will highlight efforts to develop new diagnostics and new treatments.
Revealing Hormone Links to Cancer
Hormones fuel some types of cancer, including breast cancer, in which malignant cells feed on estrogen – the principal female sex hormone. That suggests strategies to stop the cancer’s spread might include blocking the site where estrogen binds to its receptor or inhibiting the gene that controls production of estrogen.
Marion Sewer, an associate professor in the Georgia Tech School of Biology, focuses her research on a protein called liver receptor homolog 1 (LRH1). Inhibiting LRH1 could potentially slow the progression of hormone-sensitive breast cancer by stopping estrogen production.
“We are investigating LRH1 because it binds to sequences of DNA and activates the gene that produces estrogen,” says Sewer. “Although LRH1 is not typically present in normal breast tissue, it is present at high levels in breast cancer cells. We’re trying to figure out what activates it and causes it to be overproduced in cancerous tissue.”
Biologists know that activation of LRH1 cannot occur until a particular small molecule binds to it, but they are unclear about the identity of that molecule. Sewer and Eric Ortlund, an assistant professor in the Department of Biochemistry at Emory University, are trying to identify the molecule – called a ligand – that binds to LRH1 and activates it.
In collaboration with Alfred Merrill, a professor in the Georgia Tech School of Biology and the Smithgall Chair in Molecular Cell Biology, Sewer is also isolating LRH1 from breast cancer cells and using mass spectrometry techniques to identify any ligands that are bound to the receptor.
“If we can figure out what the ligand is and design some analog of the ligand that would inhibit its ability to bind to DNA and produce estrogen, we may discover a better anti-cancer therapy method,” explains Sewer.
In a related project, Sewer is investigating the genes that control production of vitamin D. These are members of the same family of genes – called cytochromes P450 – that control production of estrogen.
“Since vitamin D levels have been shown to be much lower in breast cancer patients and studies have linked insufficient vitamin D to an increased risk for breast cancer, we are investigating sphingolipid molecules that increase the presence of the genes that produce the active form of vitamin D,” notes Sewer.
The molecules – called 1-deoxysphinganines – were originally isolated from mollusks and have been shown to have anti-cancer properties. Graduate student Tenzing Phanthok and undergraduate student Viniya Patidar are investigating the role of 1-deoxysphinganine in increasing vitamin D production and in turn preventing cancer cells from multiplying.
Since these 1-deoxysphinganines are naturally produced in the body, it is possible that cancer progression is a result of altered production of 1-deoxysphinganine. Perhaps the level of these molecules could be used as a cancer biomarker or indicator, Sewer says.
Sewer’s work is funded by the National Institutes of Health, National Science Foundation and a Georgia Cancer Coalition Distinguished Scientist Award.
This project was supported by Award No. R01GM073241 from the National Institute of General Medicine Sciences (NIGMS) and Award No. MCB-0347682 from the National Science Foundation (NSF). Any opinions, findings, conclusions or recommendations expressed are those of the researcher and do not necessarily reflect the views of the NIGMS, the National Institutes of Health or the NSF.
Investigating the Role of Epigenetics in Cancer
While many biologists investigate cancer genetics – mutations in DNA sequences that cause the disease – a growing group of biologists is examining the role of cancer epigenetics, which are changes that contribute to malignancy without causing changes in DNA sequences.
Yuhong Fan, an assistant professor in Georgia Tech’s School of Biology, believes that the scientific field of epigenetics may help shape the future of cancer diagnosis and treatment.
“Cancer cells have drastically different epigenetic patterns compared to normal cells,” explains Fan, who is also a Georgia Cancer Coalition Distinguished Cancer Scholar. “Many epigenetic changes may appear prior to the development of invasive cancer, so I think that doctors might one day be able to detect epigenetic markers for cancer before a tumor appears.”
Epigenetic studies concentrate on the way the genome is marked and packaged inside a cell’s nucleus. Much of Fan’s research focuses on the role of H1 linker histones, a family of 10 proteins that helps to package the DNA within chromosomes.
Fan and Arthur Skoultchi, chair of the Department of Cell Biology at the Albert Einstein College of Medicine at New York’s Yeshiva University, previously observed the effects of partially reducing H1 levels in mice. The work showed that H1 histones are important to an organism’s normal development. Expanding on these findings, Fan recently teamed with John McDonald, chief scientist of the Ovarian Cancer Institute and associate dean for biology development in the School of Biology, to determine if the multiple H1 subtypes are regulated differently in benign and malignant ovarian cancer tissues.
“We found that some of the H1 subtypes were expressed at significantly higher levels in the cancerous tissue compared to the benign tissue and some were expressed at significantly lower levels,” notes Fan. “The most remarkable finding was that these differences, whether increases or decreases, were consistent among multiple samples.”
With this knowledge, Fan’s next step is to find out what genes and functions are affected by changes in expression of each subtype. To do this, her group plans to change the level of each H1 subtype in cancer cell culture and monitor what happens to cell growth and cell fate.
“We hope that measuring the expression level of one or more of these H1 subtypes can be used as an epigenetic biomarker for the cancer diagnosis of the future,” adds Fan. “Since the expression patterns are consistent, you could easily measure a few epigenetic characteristics, rather than looking at thousands of genes.”
Funding for Fan’s research is provided by the National Institutes of Health and the Georgia Cancer Coalition.
Examining How Ovarian Cancer Develops
Unlike many cancer biology researchers who investigate general processes underlying many cancers, John McDonald focuses his investigations broadly on one type of cancer – ovarian.
Ovarian cancer is the most lethal gynecological cancer, with the American Cancer Society predicting that in the United States alone each year, more than 20,000 women will be diagnosed with ovarian cancer and 16,000 will die from it.
“Ovarian cancer is called the silent killer because by the time symptoms arise and it’s detected, it has typically spread throughout the body,” says McDonald, chief scientist of the Ovarian Cancer Institute and associate dean for biology development in the School of Biology. “Our laboratory takes an integrated approach to studying ovarian cancer by investigating its causes, establishing accurate and reliable diagnostic tests, and developing novel and effective therapies.”
One focus of McDonald’s research is to determine how cancer cells develop in the ovaries. While it is estimated that up to 90 percent of ovarian carcinomas are derived from ovarian surface epithelial cells – cells that create the thin layer of tissue that covers the ovaries – the behavior of these cells differs from other epithelial-derived carcinomas because they become more specialized as malignancy progresses.
To investigate this behavior in more detail, McDonald and Nathan Bowen, a research scientist and Georgia Cancer Coalition Distinguished Cancer Scholar, compared the gene expression profiles of ovarian surface epithelial cells isolated from the surface of healthy ovaries with those of malignant ovarian tumors collected by the Ovarian Cancer Institute.
The results showed that more than 2,000 genes were expressed at significantly different levels in the two sample types. Genes associated with adult stem cell maintenance were expressed at a much higher level in the cells isolated from healthy ovaries.
“We found that changes in the expression of genes involved in maintaining the inertness and stem cell nature of epithelial surface ovarian cells may be instrumental in the initiation and development of ovarian cancer,” explains McDonald.
The results also showed that the surface of the ovary exhibits the characteristics of an adult stem cell niche, which is a protected environment where stem cells remain inactive until a signal triggers their cell cycle and they differentiate.
Expanding on these results, McDonald, Bowen and postdoctoral fellows Roman Mezencev and Lijuan Wang are currently examining the sensitivity of ovarian cancer stem cells and differentiated cancer cells to existing chemotherapy agents.
“The preliminary results indicate that existing chemotherapy agents may effectively kill cancer cells but not touch these cancer stem cells, which could be why ovarian tumors and other cancers frequently recur,” adds McDonald.
This work was supported by the Ovarian Cancer Institute, Georgia Cancer Coalition, Golfers Against Cancer Foundation, Ovarian Cycle Foundation, Robinson Family Foundation and Deborah Nash Harris Foundation.
Investigating DNA Repair Mechanisms
Exposure to environmental carcinogens such as tobacco smoke and ultraviolet radiation can result in various types of DNA damage and subsequently lead to the development of cancer if the damage is not repaired.
Double-strand breaks, in which both strands in the DNA double helix are severed, are particularly hazardous to cells because they can lead to genome rearrangements. And their repair is intrinsically more difficult.
Biologists typically believed that double-strand breaks could only be repaired by homologous intact DNA – until recently, when Francesca Storici, an assistant professor in Georgia Tech’s School of Biology, showed that RNA could be used as a template to directly repair DNA in yeast cells. This contradicted the dogma that genetic information had to flow from DNA to RNA.
“Using RNA that naturally resides inside a cell to repair damaged DNA could represent an additional line of defense against DNA damage,” says Storici, who is also a Georgia Cancer Coalition Scholar. “The capacity of RNA to record itself into DNA could be the basis of a wholly unexplored process of RNA-driven DNA evolution.”
These unique RNA functions may have important implications in gene targeting and gene therapy because RNA molecules mimicking RNA oligonucleotides could be generated directly in the nucleus of targeted cells via transcription from vectors.
Since her initial discovery in yeast, Storici has used RNA to repair broken chromosomal DNA in human cells in culture and to correct a base defect in the genome of bacterial cells, suggesting that RNA-templated DNA repair is a more general mechanism. She is currently examining exactly how this direct transfer of RNA information to DNA occurs.
“While we can gain a lot of insight from understanding how a cell can repair its DNA, we can also use that information to create a better method for correcting genetic defects,” notes Storici.
Her goal is to develop a tool to correct a particular mutation on a specific chromosome while causing minimal damage to the DNA. One way to do that, Storici says, might be to search for factors that facilitate delivery of the targeting molecule to the nucleus and promote the exchange of DNA strands.
To test the tool she develops, Storici is working with and constructing different human cell lines, and monitoring the repair of specific genetic defects with a simple flow cytometry assay.
Given the ability of RNA to transfer genetic information to chromosomal DNA and the possibility of amplifying RNA within cells at will, Storici plans to continue investigating new directions in gene targeting and treatment of cancer and other genetic diseases.
Understanding the Role of Sphingolipids in Cancer Development
For almost 30 years, Georgia Tech professor Alfred Merrill has been studying lipids – the fats, oils, cholesterols and certain vitamins that our bodies need to grow and survive. Today, his expertise lies in a subgroup of lipids called sphingolipids, which influence cell structure, signaling and interaction.
“The lipid backbones of sphingolipids are important cell-signaling molecules that turn on and turn off intracellular proteins that are involved in cell growth, death, and an interesting process called autophagy that has recently gained much attention in the cancer research field,” says Merrill, who is also the School of Biology’s Smithgall Chair in Molecular Cell Biology.
Autophagy – meaning “self-eating” – involves the degradation of cellular compartments, called organelles, and cellular proteins. During this process, a cell forms a vesicle that encapsulates its cytoplasm and some of the organelles and then fuses with digestive enzymes that degrade the contents of the vesicle and make them available for cell nutrition.
Interestingly, autophagy has been implicated in both cancer cell death and survival. Since Merrill’s research has shown that sphingolipid signaling is essential for creating autophagy vesicles, these metabolites may be involved in both promoting and limiting tumor growth.
Autophagy promotes cancer cell survival by allowing cells to respond to changing environmental conditions, such as nutrient deprivation. During starvation, autophagy allows cells to degrade proteins and organelles and thus obtain a source of nutrients that would not be available otherwise.
“Cancer cells use autophagy because as they are developing they have a period in which they go into a nutrient crisis because they haven’t established their own blood and nutrient flow, so they use autophagy as a way to survive in the meantime,” explains Merrill.
However, this same process of gaining nutrients can lead to tumor cell death as well. Merrill’s laboratory found that a number of anti-cancer agents promote the formation of these vesicles through sphingolipid signaling.
“Preliminary data supports the theory that the autophagic vesicles in cancer cells are unstable, so if one of their components— the sphingolipids—is out of balance, this can cause them to break apart and spill out their toxic contents, killing the cancer cell,” adds Merrill.
While the mechanism through which autophagy inhibits tumor development is still unclear, graduate student Kacee Sims is examining the role of sphingolipid pathways in the conversion of autophagy from a cancer cell survival pathway to a cell death pathway.
The project described was supported by Award No. U54GM069338 from the National Institute of General Medicine Sciences (NIGMS). Any opinions, findings, conclusions or recommendations expressed are those of the researcher and do not necessarily reflect the views of the NIGMS or the National Institutes of Health. Significant funding to support this research was also provided by the Smithgall Endowment to Georgia Tech.
Investigating the Complexity of Chromosome Breaks
Everyone has fragile sites on their chromosomes that are particularly prone to breaking, making them hot spots for rearrangements that can lead to hereditary diseases and cancer. Georgia Tech School of Biology associate professor Kirill Lobachev is trying to understand what’s special about these regions, the consequences of the breaks, and the pathways that are involved in promoting and repairing these breaks.
“It is becoming clear that the fragile sites often contain unstable repetitive sequences that can adopt unusual DNA structures,” says Lobachev. “We think that everyone is probably a carrier of these unstable motifs that can cause chromosomes to break anytime, so we ultimately want to be able to predict where a chromosome is going to break and how frequently this break will occur, and determine if we can prevent it.”
Determining whether a particular chromosomal region is predisposed to breakage requires knowledge about the structural parameters of the unstable sequences that make chromosomes fragile, such as their size or composition of the genetic sequences they contain. Using the yeast Saccharomyces cerevisiae as a model organism, Lobachev’s laboratory has been able to mimic some of the structural instability that cancer cell chromosomes exhibit.
In a recent study, Lobachev and colleagues demonstrated that DNA replication machinery sometimes stalls when it reaches a long sequence of palindromes – sequences that read the same way backward and forward. Further analysis has shown that chromosomes break when DNA replication is slowed or altered.
“Long palindromes were known to change the shape of DNA from a double helix into a hairpin or cruciform structure, but this was one of the first studies to show that these changes could affect DNA integrity,” explains Lobachev.
In addition, Lobachev and postdoctoral fellow Vidhya Narayanan determined that palindromic sequences induce a particular type of DNA break that is a precursor to a process involved in cancer called gene amplification. Amplification of genes involved in metabolism or inactivation of drugs can lead to chemotherapy resistance, and amplification of genes that turn normal cells into cancer cells are known to occur in several late-stage cancers.
They showed that gene amplification depends on the location of an oncogene relative to the break – called a hairpin-capped double strand break – and the end of the chromosome. The study indicated that restricting breakage of the unstable sequences may be a promising strategy for pharmaceutical cancer prevention and treatment.
In the future, knowing what genetic sequences are more likely to lead to chromosomal fragility and being able to explore genetic pathways involved in this process may help researchers identify persons who might be prone to developing cancer, adds Lobachev.
Collaboration with the Georgia Cancer Coalition
The Georgia Cancer Coalition’s (GCC) mission is to reduce the number of cancer deaths in Georgia. One key initiative toward accomplishing that goal is naming Georgia Cancer Coalition Distinguished Cancer Clinicians and Scientists. In concert with Georgia’s academic universities, the GCC supports the recruitment of national leaders in cancer research to Georgia.
At Georgia Tech, 11 researchers have been named Distinguished Cancer Scholars, including:
- Ravi Bellamkonda, professor, biomedical engineering
- Nathan Bowen, senior research scientist, biology
- Erin Dickerson, research scientist, biology
- Yuhong Fan, assistant professor, biology
- Melissa Kemp, assistant professor, biomedical engineering
- Valeria Tohver Milam, assistant professor, materials science and engineering
- Shuming Nie, professor, biomedical engineering
- Marion Sewer, associate professor, biology
- Francesa Storici, assistant professor, biology
- Dongmei “May” Wang, assistant professor, biomedical engineering
- Ming Yuan, assistant professor, industrial and systems engineering
The Georgia Cancer Coalition has also awarded seven Cancer Research Awards to Georgia Tech faculty members investigating how to prevent, treat and cure breast, ovarian and prostate cancers. Michelle Dawson, an assistant professor in Georgia Tech’s School of Chemical and Biomolecular Engineering, recently received one of these grants for her research into the development of specialized cells designed as gene delivery vehicles to target and treat breast cancer.
Groups
Status
- Workflow status: Published
- Created by: Claire Labanz
- Created: 11/05/2014
- Modified By: Fletcher Moore
- Modified: 10/07/2016
Categories
Keywords
User Data