news
Bringing structure to bear on RNA viruses: SoB scientists provide new insight on viral packing
Primary tabs
Yingying Zeng, a graduate student in the School of Biology, is the lead author on a new paper that describes the complete structure of satellite tobacco mosaic virus (STMV). This is the first model for the structure of any virus that specifies the position of every single atom. Ms. Zeng combined high-resolution data from x-ray crystallography, chemical data on the structure of the RNA genome, and knowledge-based molecular modeling methods to develop her model. STMV is a small virus that has served for many years as a model system for investigating the relationships between viral structure and function. The new model has implications for understanding the pathway of viral assembly. These methods can be extended to investigate the structures of human viral pathogens, and, in the long run, to the design of novel drugs aimed at inhibiting viral assembly.
This was a collaborative effort headed by Dr. Steve Harvey (Georgia Tech School of Biology), and it included contributions from Dr. Christine Heitsch (Georgia Tech School of Mathematics) and Drs. Steven Larson and Alexander McPherson (University of California, Irvine). The paper will soon appear in Journal of Structural Biology.
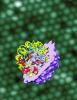
Caption for figure: The picture shows the virus structure with half of the outer shell (the viral capsid, lavender) cut away to reveal the RNA genome on the inside. Only the RNA backbone is shown in the image. The red RNA double helices lie around an axis of five-fold symmetry just inside the capsid. Other RNA double helices that lie just inside the capsid are colored yellow, while the remainder of the RNA backbone is shown in pale blue. The background image is an electron micrograph of one face of the crystal from which Drs. Larson and McPherson determined the x-ray structure of the virus.
Status
- Workflow status: Published
- Created by: Troy Hilley
- Created: 08/20/2012
- Modified By: Fletcher Moore
- Modified: 10/07/2016
Categories
User Data